
MGI's DNBSEQ-T7+ platform has successfully completed comprehensive verification with Olink ™ Explore HT, the industry's leading high-throughput proteomics solution. This milestone extends the versatility of MGI's sequencing ecosystem into next-generation proteomics, offering researchers a powerful, validated workflow for large-scale protein biomarker discovery
MGI has previously verified the sequencing workflows for different Olink technology libraries on the T7, T1+, and G400 platforms, with data output meeting expectations. This means that customers with different MGI sequencing platforms can now perform high-quality proteomic library sequencing, adding one more option for multi-omics research.
Olink Panel | MGI Sequencing Platform |
Olink™ Explore HT | T7+ & T7 |
Olink™ Reveal | T1+ & G400 |
Verification at a Glance
The joint validation study systematically evaluated Olink™ Explore HT performance on DNBSEQ-T7 and T7+ platforms, benchmarking against established sequencing workflows. The evaluation focused on the consistency of performance, quantitative correlations, assay variability, and detectability.
Result
1) Read yield of the Olink™ Explore HT library on the MGI platform meets expectations and is comparable to the existing Olink technology sequencing workflow: 1 T7+ FCL or 2 T7 FCL can generate the required data volume for Explore HT analysis.
2) Data generated from MGI platforms demonstrates high consistency with that from other platforms, while also showing strong intra-platform reproducibility.
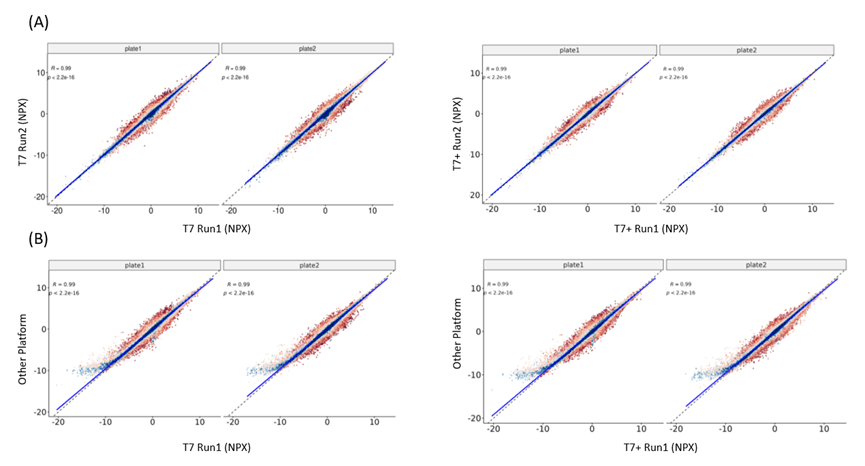
(A) Pearson correlation between runs on MGI Sequencing Platform (B) Pearson correlation between T7+/ T7 and other sequencing platform
3) The CV and detectability are comparable across sequencing platforms, ensure reliable protein detection and robust biological insights.
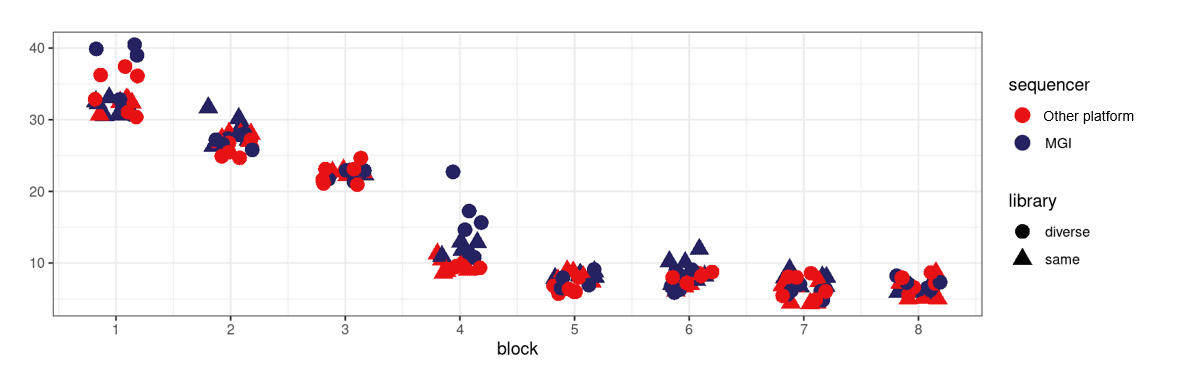
NPX-based intra CV of Explore HT library in T7+ sequencing platform
Customer Cases
Based on the aforementioned validation workflow, MGI customers also completed the first sequencing run of Olink™ Explore HT real-sample libraries on the T7+ platform, with parallel comparison to other validated platforms. This test included 48 Concordance test plasma samples, all of which passed Olink™ Concordance Test. Detailed results are summarized as follows:
Term | Result |
QC warning | PASS |
CV | PASS |
Correlation | PASS (Mean=0.86; Median=0.93) |
Regression | PASS (0.92) |
Concordance Test | PASS |
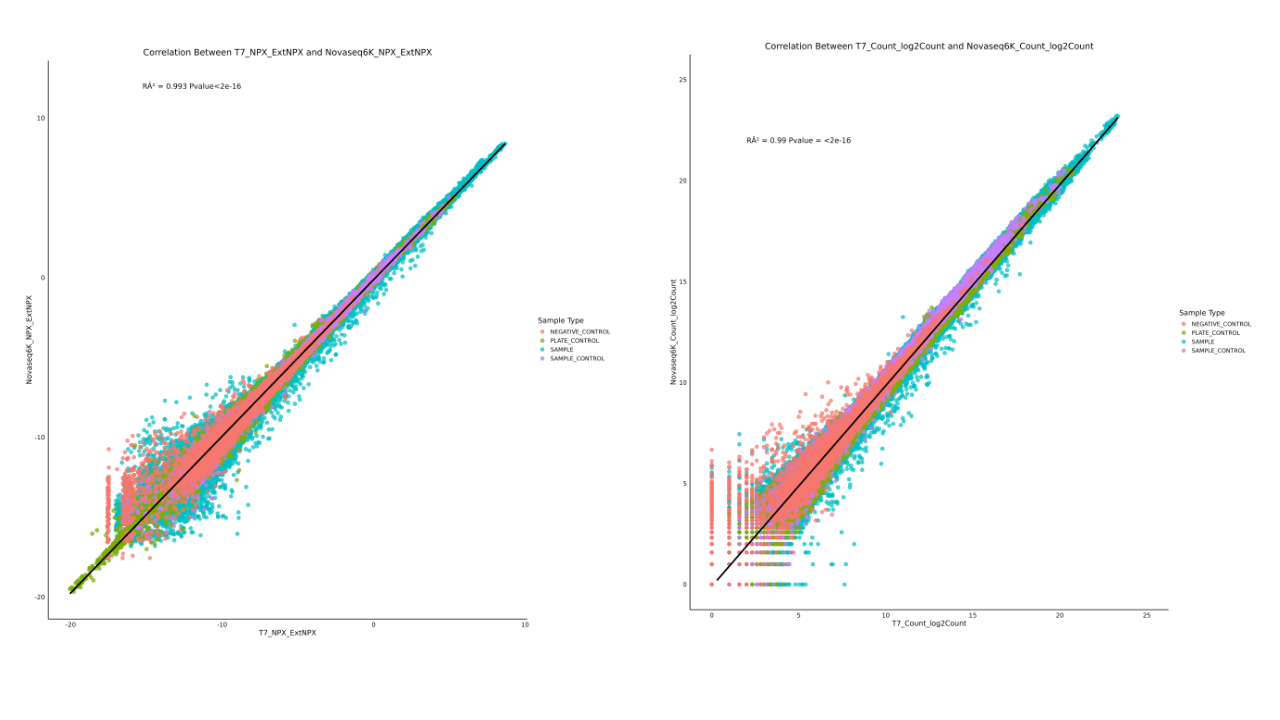
The correlation coefficients for sample Counts (A) and NPX values (B) between the T7+ platform and other platforms are all above 0.99.
The above test results demonstrate that Olink™ Explore HT performs excellently on MGI's latest high-throughput sequencing platform, the T7+, delivering high‑quality data and outcomes. This confirms that MGI T7+ provides researchers with a high-throughput proteomics sequencing solution.
Why DNBSEQ-T7+ for High-Throughput Proteomics?
As MGI's flagship DNBSEQ™ platform celebrating a decade of innovation, the T7+ seamlessly integrates core DNBSEQ™ technology with the StandardMPS 2.0 sequencing chemistry, redefining the boundaries of large-scale sequencing efficiency through its quantity and speed, quality and cost-effectiveness performance.
Supporting flexible loading of 4 flow cells with up to 3.6 Tb output per cell, the T7+ delivers over 14 Tb of high-quality data in just 24 hours in PE150 mode. Designed with intelligence and user-centricity in mind, its compact ~1 m² footprint, 30% reagent reduction, and 50% waste reduction not only minimize operational costs and space requirements but also position it as a powerful yet "warm-hearted" research partner, providing a solid, reliable hardware foundation for diverse applications including large-scale population genomics, deep multi-omics analysis, and clinical-grade sequencing.
Get Started
The T7+ platform is now validated and ready for Olink™ Explore HT workflows. Contact your MGI representative to discuss integration into your proteomics pipeline, or visit to learn more about our expanding application portfolio.
biomarkers
proteomics
NGS
Share






