DNBSEQ Platforms: Superior Performance in Cross-Platform Benchmarking Study Demonstrated
Apr 14, 2026
Share article

A comprehensive whole-genome sequencing (WGS) benchmarking study has evaluated MGI's DNBSEQ platforms against three leading short-read sequencing systems, the Illumina NovaSeq 6000, Element AVITI and PacBio Onso, using the NIST GIAB reference sample HG002/NA24385 at a uniform 30× effective sequencing depth. Across all evaluated metrics, raw data quality, alignment fidelity and variant calling accuracy, DNBSEQ platforms demonstrated consistent superiority, with SNP detection reaching an F-measure of 99.80% and Clean Q30 scores exceeding 98%.
Study design and platform overview
Three DNBSEQ instruments were evaluated: the DNBSEQ-T7+, DNBSEQ-T1+ and DNBSEQ-T7. These were benchmarked against PCR-free WGS datasets generated on the Illumina NovaSeq 6000, Element AVITI and PacBio Onso. All datasets were subsampled to identical effective sequencing depth using seqtk (v1.2) to ensure a balanced, fair comparison. Quality control was performed with SOAPnuke (v2.1.9), including adapter trimming, low-quality read filtering and removal of high-N content reads.
Alignment was carried out against the GRCh38 reference genome using FPGA-accelerated MegaBOLT software implementing the BWA-MEM2 algorithm. Performance in complex genomic regions was assessed using NIST genome stratification intervals. Variant calling used the MGI PanVariants pipeline for DNBSEQ data and DeepVariant (v1.9.0) for comparator platforms, evaluated against the NIST Reference Material v4.2.1 benchmark set using RTG Tools vcfeval, in accordance with GA4GH-recommended methods.
Raw data quality
DNBSEQ platforms achieved Clean Q30 scores above 98% across all three instruments, with minimal inter-platform variability. Comparator platforms ranged from 91% to 99%, reflecting greater inconsistency. DNBSEQ also required only 93 Gbp of raw data to achieve 30× effective coverage, demonstrating more efficient data utilisation than competing systems. Q40 scores exceeded 94%, and internal testing reported Q50 scores above 80%.
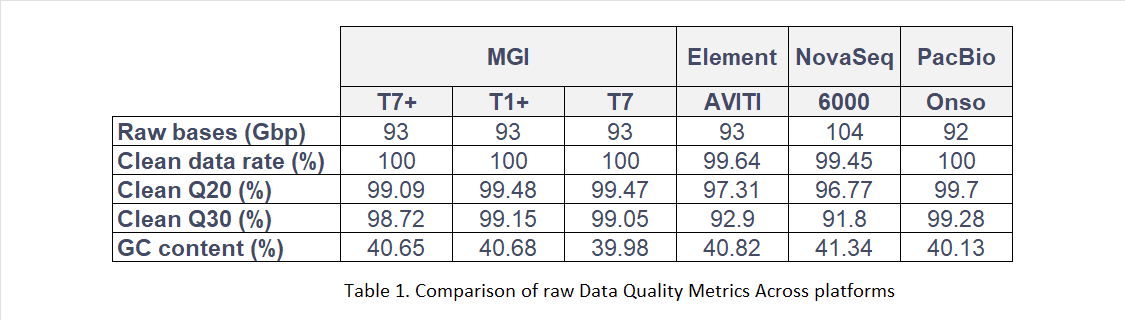
Due to differences in sequencing throughput across platforms, uniform subsampling was applied to FASTQ files to ensure comparability. Each dataset was subsampled to 30× coverage using seqtk (v1.2). Quality control was then performed using SOAPnuke (v2.1.9), including adapter trimming, low-quality filtering and removal of reads with high N content.
Alignment metrics at 30× effective depth
At 30× effective sequencing depth, DNBSEQ platforms matched or outperformed all comparators across four key alignment metrics.
Duplicate rate: DNBSEQ maintained significantly lower duplication levels, preserving effective data yield across all three instruments.
Mismatch rate: DNBSEQ achieved a mismatch rate of 0.34%, compared to over 0.5% for both the Illumina NovaSeq 6000 and PacBio Onso, the lowest error rate across all tested platforms.
Coverage uniformity (≥20×): All DNBSEQ platforms exceeded 93% of the genome covered at ≥20×. PacBio Onso reached 88% under equivalent conditions.
Homopolymer region coverage (≥10×): DNBSEQ maintained >98% coverage in homopolymer regions, compared to 93% for PacBio Onso, a particularly relevant metric for clinical and population genomics applications where repeat-rich regions are diagnostically important.
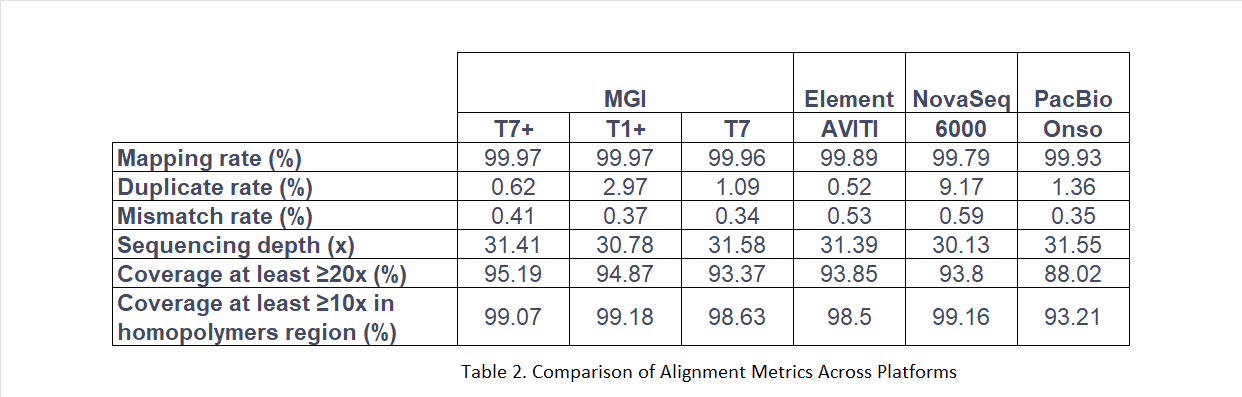
Following subsampling and quality control, alignment was performed using the FPGA-accelerated MegaBOLT software, implementing the BWA-MEM2 algorithm against the GRCh38 reference genome. Genome stratification intervals published by NIST were used to evaluate performance in complex genomic regions.
Variant calling accuracy
SNP detection: DNBSEQ achieved a precision of 99.80%, sensitivity of 99.78% and an F-measure of 99.80%, outperforming Element AVITI (F-measure: 99.60%), Illumina NovaSeq 6000 (99.57%) and PacBio Onso (98.93%).
Indel detection: DNBSEQ reached an indel F-measure exceeding 99.55%, comparable to Element AVITI (99.52%) and higher than Illumina NovaSeq 6000 (99.42%) and PacBio Onso (98.38%).
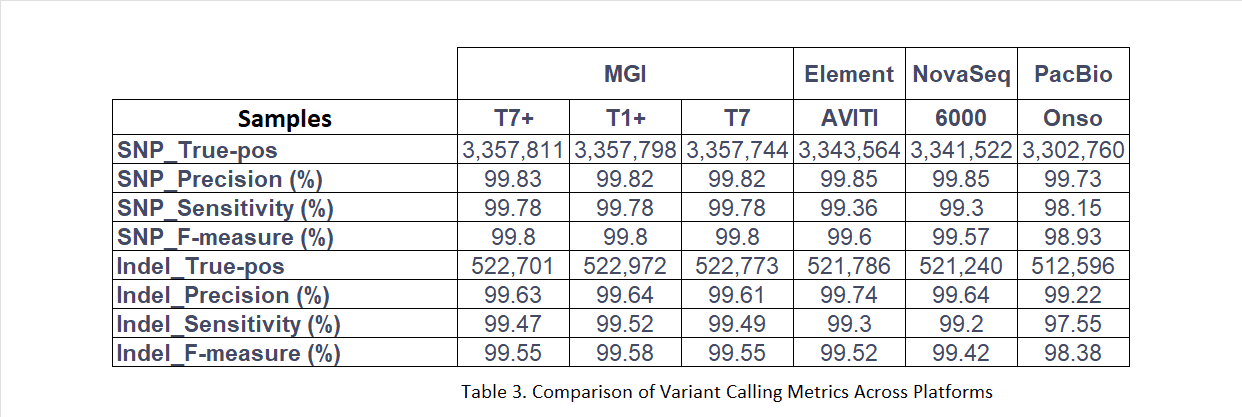
SNP and indel detection was conducted using the PanVariants pipeline for DNBSEQ data and DeepVariant (v1.9.0) for comparator platforms. Performance was evaluated using RTG Tools against the NIST v4.2.1 benchmark set.
Metric used:
Precision = TP / (TP + FP)
Sensitivity = TP / (TP + FN)
F-measure = 2 × Precision × Sensitivity / (Precision + Sensitivity)
Frequently asked questions
How does DNBSEQ compare to Illumina NovaSeq 6000 for SNP detection?
In this study, DNBSEQ achieved an SNP F-measure of 99.80% versus 99.57% for the Illumina NovaSeq 6000, using DeepVariant v1.9.0 for NovaSeq and the MGI PanVariants pipeline for DNBSEQ, both evaluated against NIST GIAB v4.2.1.
What reference sample was used in this WGS benchmarking study?
The study used HG002/NA24385, a globally recognised NIST GIAB reference material, at 30× effective sequencing depth with uniform subsampling applied to FASTQ files via seqtk v1.2.
What is the mismatch rate of DNBSEQ platforms?
DNBSEQ platforms achieved a mismatch rate of 0.34% at 30× effective depth, the lowest of all platforms tested, compared to over 0.5% for the Illumina NovaSeq 6000 and PacBio Onso.
How does DNBSEQ perform in homopolymer regions?
DNBSEQ maintained >98% coverage at ≥10× depth in homopolymer regions, compared to 93% for PacBio Onso, indicating better uniformity in genomic regions historically challenging for short-read sequencing.
What variant calling tools were used?
The MGI PanVariants pipeline was used for DNBSEQ datasets; DeepVariant v1.9.0 was used for all comparator platforms. Benchmarking was conducted using RTG Tools vcfeval against the NIST v4.2.1 truth set, following GA4GH-recommended best practices.
References:
Benchmarking challenging small variants with linked and long reads. Cell Genom. 2022 May;2(5):100128. doi: 10.1016/j.xgen.2022.100128. PMID: 36452119; PMCID: PMC9706577.
Comparing Variant Call Files for Performance Benchmarking of Next-Generation Sequencing Variant Calling Pipelines. bioRxiv 023754; doi: https://doi.org/10.1101/023754.
Best practices for benchmarking germline small-variant calls in human genomes. Nat Biotechnol. 2019 May;37(5):555-560. doi: 10.1038/s41587-019-0054-x. Epub 2019 Mar 11. Erratum in: Nat Biotechnol. 2019 May;37(5):567. doi: 10.1038/s41587-019-0108-0. PMID: 30858580; PMCID: PMC6699627.
A universal SNP and small-indel variant caller using deep neural networks. Nature Biotechnology 36, 983–987 (2018). . doi: https://doi.org/10.1038/nbt.4235. Shen W, Le S, Li Y, Hu F. SeqKit: A Cross-Platform and Ultrafast Toolkit for FASTA/Q File Manipulation. PLoS One. 2016 Oct 5;11(10):e0163962. doi: 10.1371/journal.pone.0163962. PMID: 27706213; PMCID: PMC5051824.
SOAPnuke: a MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. Gigascience. 2018 Jan 1;7(1):1-6. doi: 10.1093/gigascience/gix120. PMID: 29220494; PMCID: PMC5788068.
Efficient Architecture-Aware Acceleration of BWA-MEM for Multicore Systems. IEEE Parallel and Distributed Processing Symposium (IPDPS), 2019. 10.1109/IPDPS.2019.00041.
The GIAB genomic stratifications resource for human reference genomes. Nat Commun 15, 9029 (2024). https://doi.org/10.1038/s41467
Share this article :
Share




